Core Functionality#
otoole’s functionality includes converting between data formats, processing
solution files, and visualising the model. These core functions, which provide a method
to turbo-charge your modelling, are described on this page!
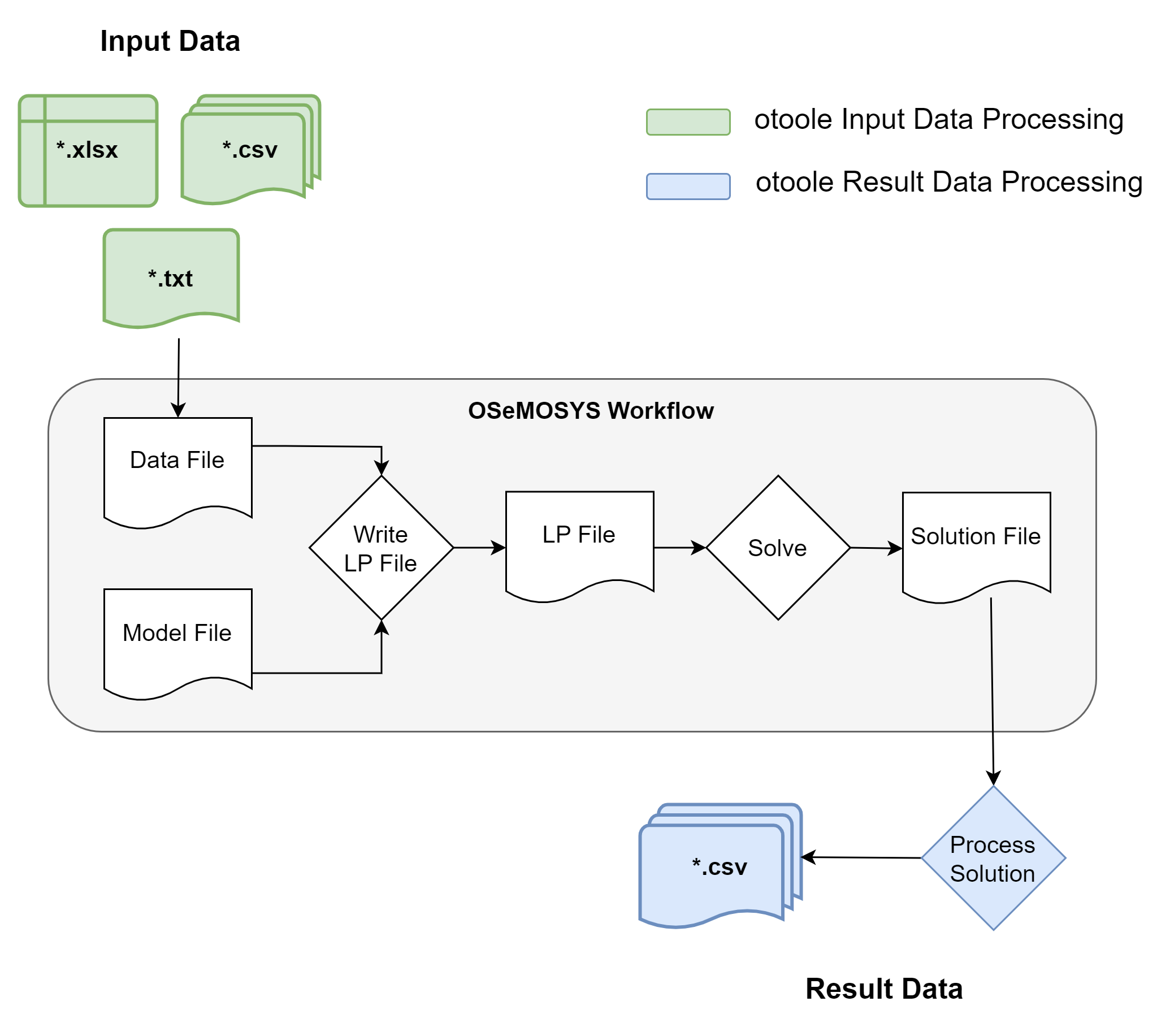
As shown in the diagram, otoole deals primarily with data before and after OSeMOSYS.
Data work prior to the generation of a datafile, which is read in by GLPK, is called
pre-processing. Anything that happens with the immediate outputs of a solver, such as
the recommended open-source solvers GLPK, CBC, HiGHS, or the commercial solvers CPLEX and
Gurobi, is called results and post-processing.
Note
While otoole is targeted at OSeMOSYS users, the functionality can easily be extended
to work with any workflow that involves the use of a MathProg file!
Data Conversion#
Overview#
otoole supports different data pre-processing conversions so as to ease the tasks of
the OSeMOSYS modeller. The modeller can generate data in any one of the formats and
convert it to another format through a convert command. otoole currently supports
conversion between the following formats:
Excel
A folder of CSV files
GNU MathProg datafile
otoole convert#
The otoole convert` command allows you to convert between various different
input formats:
$ otoole convert --help
usage: otoole convert [-h] [--write_defaults] {csv,datafile,excel} {csv,datafile,excel} from_path to_path config
positional arguments:
{csv,datafile,excel} Input data format to convert from
{csv,datafile,excel} Input data format to convert to
from_path Path to file or folder to convert from
to_path Path to file or folder to convert to
config Path to config YAML file
optional arguments:
-h, --help show this help message and exit
--write_defaults Writes default values
--keep_whitespace Keeps leading/trailing whitespace
Deprecated since version v1.0.0: The datapackage format is no longer supported
New in version v1.0.0: The config positional argument is now required
Result Processing#
Overview#
With small OSeMOSYS models, it is normally fine to use the free open-source GLPK solver. If you do, then OSeMOSYS will write out a full set of results as a folder of CSV files. As you progress to larger models, the performance constraints of GLPK quickly become apparent. CBC and HiGHS are alternative open-source solvers which offer better performance than GLPK and can handle much larger models.
otoole currently supports using GLPK, CBC, HiGHS, CPLEX or Gurobi with all versions of
GNU MathProg OSeMOSYS - the long, short and fast versions.
The long version includes all results as variables within the formulation, so the
otoole results command parses the solution file, extracts the required variables,
and produces a folder of CSV files containing the results in an identical format
to if you had used GLPK.
The short and fast versions omit a large number of these calculated result variables so as to speed up the model matrix generation and solution times.
otoole results#
The results command creates a folder of CSV result files from a GLPK, CBC, CLP,
Gurobi or CPLEX solution file together with the input data:
$ otoole results --help
usage: otoole results [-h] [--glpk_model GLPK_MODEL] [--write_defaults]
{cbc,cplex,glpk,gurobi,highs} {csv} from_path to_path {csv,datafile,excel} input_path config
positional arguments:
{cbc,cplex,glpk,gurobi,highs} Result data format to convert from
{csv} Result data format to convert to
from_path Path to solver solution file
to_path Path to file or folder to convert to
{csv,datafile,excel} Input data format
input_path Path to input_data
config Path to config YAML file
options:
-h, --help show this help message and exit
--glpk_model GLPK_MODEL GLPK model file required for processing GLPK results
--write_defaults Writes default values
New in version v1.0.0: The config positional argument is now required
New in version v1.1.0: The input_data_format and input_path positional arguments are now required
supporting any supported format of input data for results processing.
Deprecated since version v1.0.0: The --input_datapackage flag is no longer supported
Deprecated since version v1.1.0: The --input_datapackage and --input_datafile flags
have been replaced by new positional arguments input_data_format and input_path
Setup#
The setup module in otoole allows you to generate template files to
quickly get up and running.
otoole setup#
The setup command allows you to generate a template user configuration file,
useful for conversion and result commands, and template input csv
data:
$ otoole setup --help
usage: otoole setup [-h] [--write_defaults] [--overwrite] {config,csv} data_path
positional arguments:
{config,csv} Type of file to setup
data_path Path to file or folder to save to
optional arguments:
-h, --help show this help message and exit
--write_defaults Writes default values
--overwrite Overwrites existing data
Warning
The template files are generated based on a specific version of OSeMOSYS, users will need to adapt the template data for their own needs
Visualization#
Overview#
The visualization module in otoole allows you to visualise the reference energy system.
(with more visualisations to come!)
otoole viz#
The viz command allows you to visualise aspects of the model. Currently, only
visualising the reference energy system through the vis res command is supported:
$ otoole viz res --help
usage: otoole viz res [-h] {csv,datafile,excel} data_path resfile config
positional arguments:
{csv,datafile,excel} Input data format
data_path Input data path
resfile Path to reference energy system
config Path to config YAML file
optional arguments:
-h, --help show this help message and exit
Note
The resfile command should include a file ending used for images,
including bmp, jpg, pdf, png etc. The Graphviz library
used to layout the reference energy system will interpret the file ending.
Warning
If you encounter a Graphviz dependencey error, please follow Graphviz installation instructions described in the visualization examples.
Validation#
The validation module in otoole checks technology and fuel names against a
standard or user defined configuration file.
otoole validate#
The validate command allows you to identify any incorrectly named technologies
or fuels, by comparing against a user defined validation configuration file.
Moreover, otoole will check if any technology or fuel are unconnected from
the rest of the model:
$ otoole validate --help
usage: otoole validate [-h] [--validate_config VALIDATE_CONFIG] {csv,datafile,excel} data_file user_config
positional arguments:
{csv,datafile,excel} Input data format
data_file Path to the OSeMOSYS data file to validate
user_config Path to config YAML file
optional arguments:
-h, --help show this help message and exit
--validate_config VALIDATE_CONFIG
Path to a user-defined validation-config file
